Model training#
In this notebook we focus on how to train InterScale using the molecular cartography dataset used from Legnini et al., 2023 which we preprocessed in the previous tutorial.
Requirements:
preprocessed AnnData object
.yamlconfig file saved in/config_files/.
Import packages#
import scanpy as sc
from pathlib import Path
import warnings
warnings.filterwarnings("ignore")
import interscale
from interscale.config import load_config
from interscale.tl import prepare_geome_dataset, set_full_reproducibility
from interscale.geome_dataloader import GraphAnnDataModule
set_full_reproducibility()
Path.cwd().resolve().parent.parent
PosixPath('/Users/francesca.drummer/Documents/1_Projects/A3-InterScale/interscale')
from pathlib import Path
import sys
# Find repo root by locating paths.py
BASE_DIR_PROJECT = Path.cwd().resolve().parent.parent
sys.path.insert(0, str(BASE_DIR_PROJECT))
DATA = "legnini"
Load data and config#
First we load the preprocessed data from the previous tutorial.
adata = sc.read_h5ad(f"{BASE_DIR_PROJECT}/data/{DATA}_pp.h5ad")
adata
AnnData object with n_obs × n_vars = 43762 × 88
obs: 'Cell', 'Area', 'x', 'y', 'sample', 'condition', 'organoid', 'obs_names', 'split'
var: 'gene_ids', 'feature_types'
uns: 'spatial_neighbors'
obsm: 'spatial'
layers: 'log1p_norm', 'norm_ftsqrt', 'raw'
obsp: 'spatial_connectivities', 'spatial_distances'
cfg = load_config(Path(f"{BASE_DIR_PROJECT}/config_files/{DATA}_example.yaml"))
Set up model#
We need to specify:
prediction_task: Prediction task can either be classification or regressionprediction_level: Which level the predictions should be performed on: either (1) tissue label, e.i. condition (graph), (2) node label (node) for cell type or niche prediciton, or (3) GEX prediction.
Additionally, we define dataset specific keys:
prediction_obs: Label in adata.obs to be predicted. Only required for classification tasks.layer_key: Defines which GEX matrix to retrieve from adata.layersample_key: adata.obs used to split the samples into PyG Data objectsgroup_label: Optional: only if we have a adata.obs group that we want to stratify during sampling
interscale.model.CombinedModel._setup_anndata(
adata=adata, prediction_task="regression", layer_key="log1p_norm", sample_key_list=["sample"], split_key="split"
)
Anndata setup with scvi-tools version 1.4.2.
Summary Statistics ┏━━━━━━━━━━━━━━━━━━┳━━━━━━━┓ ┃ Summary Stat Key ┃ Value ┃ ┡━━━━━━━━━━━━━━━━━━╇━━━━━━━┩ │ n_sample_key_0 │ 17 │ │ n_split_key │ 3 │ │ n_x │ 88 │ └──────────────────┴───────┘
Data Registry ┏━━━━━━━━━━━━━━┳━━━━━━━━━━━━━━━━━━━━━━━━━━━━━━━━━┓ ┃ Registry Key ┃ scvi-tools Location ┃ ┡━━━━━━━━━━━━━━╇━━━━━━━━━━━━━━━━━━━━━━━━━━━━━━━━━┩ │ sample_key_0 │ adata.obs['_scvi_sample_key_0'] │ │ split_key │ adata.obs['_scvi_split_key'] │ │ x │ adata.layers['log1p_norm'] │ └──────────────┴─────────────────────────────────┘
sample_key_0 State Registry ┏━━━━━━━━━━━━━━━━━━━━━┳━━━━━━━━━━━━━┳━━━━━━━━━━━━━━━━━━━━━┓ ┃ Source Location ┃ Categories ┃ scvi-tools Encoding ┃ ┡━━━━━━━━━━━━━━━━━━━━━╇━━━━━━━━━━━━━╇━━━━━━━━━━━━━━━━━━━━━┩ │ adata.obs['sample'] │ slide1_A2-1 │ 0 │ │ │ slide1_A2-2 │ 1 │ │ │ slide1_B2-1 │ 2 │ │ │ slide1_B2-2 │ 3 │ │ │ slide1_B2-3 │ 4 │ │ │ slide1_C2-1 │ 5 │ │ │ slide1_C2-2 │ 6 │ │ │ slide1_C2-3 │ 7 │ │ │ slide1_C2-5 │ 8 │ │ │ slide1_D2-2 │ 9 │ │ │ slide1_D2-3 │ 10 │ │ │ slide4_A2-1 │ 11 │ │ │ slide4_A2-2 │ 12 │ │ │ slide4_A2-3 │ 13 │ │ │ slide4_B2-1 │ 14 │ │ │ slide4_B2-2 │ 15 │ │ │ slide4_B2-3 │ 16 │ └─────────────────────┴─────────────┴─────────────────────┘
split_key State Registry ┏━━━━━━━━━━━━━━━━━━━━┳━━━━━━━━━━━━┳━━━━━━━━━━━━━━━━━━━━━┓ ┃ Source Location ┃ Categories ┃ scvi-tools Encoding ┃ ┡━━━━━━━━━━━━━━━━━━━━╇━━━━━━━━━━━━╇━━━━━━━━━━━━━━━━━━━━━┩ │ adata.obs['split'] │ test │ 0 │ │ │ train │ 1 │ │ │ val │ 2 │ └────────────────────┴────────────┴─────────────────────┘
When run interactively, then check whether the the variables used to set up the anndata are the same as in the config.
model = interscale.model.CombinedModel(adata, cfg=cfg)
print(model._model_summary_string)
regression model for node prediction.
Dual Decoder Combined Module:
Local Module: GCN Local Component:
n_layers: 2,
n_hidden: 256,
n_embed: 16,
dropout_local: 0.1
Global Module: Transformer Encoder Global Component:
max_seq_len: 4299,
n_heads: 4,
act_func: relu,
num_layers: 2,
Dataloader#
pyg_data_list, _ = prepare_geome_dataset(adata, cfg)
dm = GraphAnnDataModule(
datas=pyg_data_list,
num_workers=1,
batch_size=int(cfg.dataset.batch_size),
pct_mask_nodes=cfg.dataset.pct_mask_nodes,
learning_type="node",
)
INFO Creating graph using `generic` coordinates and `None` transform and `1` libraries.
INFO Creating graph using `generic` coordinates and `None` transform and `1` libraries.
INFO Creating graph using `generic` coordinates and `None` transform and `1` libraries.
INFO Creating graph using `generic` coordinates and `None` transform and `1` libraries.
INFO Creating graph using `generic` coordinates and `None` transform and `1` libraries.
INFO Creating graph using `generic` coordinates and `None` transform and `1` libraries.
INFO Creating graph using `generic` coordinates and `None` transform and `1` libraries.
INFO Creating graph using `generic` coordinates and `None` transform and `1` libraries.
INFO Creating graph using `generic` coordinates and `None` transform and `1` libraries.
INFO Creating graph using `generic` coordinates and `None` transform and `1` libraries.
INFO Creating graph using `generic` coordinates and `None` transform and `1` libraries.
INFO Creating graph using `generic` coordinates and `None` transform and `1` libraries.
INFO Creating graph using `generic` coordinates and `None` transform and `1` libraries.
INFO Creating graph using `generic` coordinates and `None` transform and `1` libraries.
INFO Creating graph using `generic` coordinates and `None` transform and `1` libraries.
INFO Creating graph using `generic` coordinates and `None` transform and `1` libraries.
INFO Creating graph using `generic` coordinates and `None` transform and `1` libraries.
INFO Creating graph using `generic` coordinates and `None` transform and `1` libraries.
INFO Creating graph using `generic` coordinates and `None` transform and `1` libraries.
Training#
model.train(max_epochs=20, datamodule=dm, early_stopping=True, wandb_use=False)
Steps per epoch 4
cross-cell per gene correlation metrics
Model checkpoint will be saved in: /Users/francesca.drummer/Documents/1_Projects/A3-InterScale/results/legnini23/dual_legnini23_regr_node_44_GCN_self-attn-transformer_model.ckpt
┏━━━┳━━━━━━━━━━━━━━━┳━━━━━━━━━━━━━━━━━━━━━━━━━━━┳━━━━━━━━┳━━━━━━━┳━━━━━━━┓ ┃ ┃ Name ┃ Type ┃ Params ┃ Mode ┃ FLOPs ┃ ┡━━━╇━━━━━━━━━━━━━━━╇━━━━━━━━━━━━━━━━━━━━━━━━━━━╇━━━━━━━━╇━━━━━━━╇━━━━━━━┩ │ 0 │ module │ DualDecoderCombinedModule │ 51.2 K │ train │ 0 │ │ 1 │ loss │ SmoothL1Loss │ 0 │ train │ 0 │ │ 2 │ train_metrics │ MetricCollection │ 0 │ train │ 0 │ │ 3 │ valid_metrics │ MetricCollection │ 0 │ train │ 0 │ │ 4 │ test_metrics │ MetricCollection │ 0 │ train │ 0 │ └───┴───────────────┴───────────────────────────┴────────┴───────┴───────┘
Trainable params: 51.2 K Non-trainable params: 0 Total params: 51.2 K Total estimated model params size (MB): 0 Modules in train mode: 63 Modules in eval mode: 0 Total FLOPs: 0
┏━━━━━━━━━━━━━━━━━━━━━━━━━━━┳━━━━━━━━━━━━━━━━━━━━━━━━━━━┓ ┃ Validate metric ┃ DataLoader 0 ┃ ┡━━━━━━━━━━━━━━━━━━━━━━━━━━━╇━━━━━━━━━━━━━━━━━━━━━━━━━━━┩ │ val_cosine_similarity │ nan │ │ val_global_loss │ 1.4509000778198242 │ │ val_local_loss │ 1.397918701171875 │ │ val_loss │ 1.4244095087051392 │ │ val_mse │ 7.19042444229126 │ │ val_pearson_corr │ 0.016671981662511826 │ │ val_r2 │ -1.609514594078064 │ └───────────────────────────┴───────────────────────────┘
on_test_epoch_end 20
┏━━━━━━━━━━━━━━━━━━━━━━━━━━━┳━━━━━━━━━━━━━━━━━━━━━━━━━━━┓ ┃ Test metric ┃ DataLoader 0 ┃ ┡━━━━━━━━━━━━━━━━━━━━━━━━━━━╇━━━━━━━━━━━━━━━━━━━━━━━━━━━┩ │ test_cosine_similarity │ nan │ │ test_global_loss │ 1.6440597772598267 │ │ test_local_loss │ 1.5908509492874146 │ │ test_loss │ 1.6174554824829102 │ │ test_mse │ 8.017255783081055 │ │ test_pearson_corr │ 0.022260380908846855 │ │ test_r2 │ -4.7129316329956055 │ └───────────────────────────┴───────────────────────────┘
model.history_.plot_history(subset_term="loss")
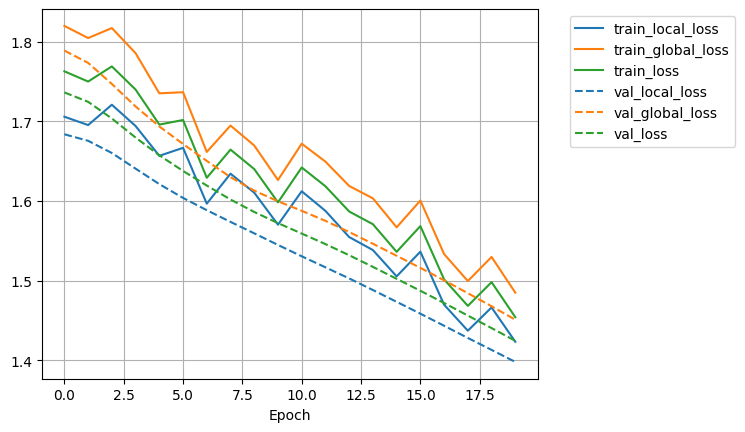
model.save(f"{BASE_DIR_PROJECT}/results/{DATA}")
Infer results#
result = model.get_model_output(adata)
Load GEX from .layers/log1p_norm
INFO Creating graph using `generic` coordinates and `None` transform and `1` libraries.
INFO Creating graph using `generic` coordinates and `None` transform and `1` libraries.
INFO Creating graph using `generic` coordinates and `None` transform and `1` libraries.
INFO Creating graph using `generic` coordinates and `None` transform and `1` libraries.
INFO Creating graph using `generic` coordinates and `None` transform and `1` libraries.
INFO Creating graph using `generic` coordinates and `None` transform and `1` libraries.
INFO Creating graph using `generic` coordinates and `None` transform and `1` libraries.
INFO Creating graph using `generic` coordinates and `None` transform and `1` libraries.
INFO Creating graph using `generic` coordinates and `None` transform and `1` libraries.
INFO Creating graph using `generic` coordinates and `None` transform and `1` libraries.
INFO Creating graph using `generic` coordinates and `None` transform and `1` libraries.
INFO Creating graph using `generic` coordinates and `None` transform and `1` libraries.
INFO Creating graph using `generic` coordinates and `None` transform and `1` libraries.
INFO Creating graph using `generic` coordinates and `None` transform and `1` libraries.
INFO Creating graph using `generic` coordinates and `None` transform and `1` libraries.
INFO Creating graph using `generic` coordinates and `None` transform and `1` libraries.
INFO Creating graph using `generic` coordinates and `None` transform and `1` libraries.
Save results#
result.write(f"{BASE_DIR_PROJECT}/data/{DATA}_trained.h5ad")
