InterScale#
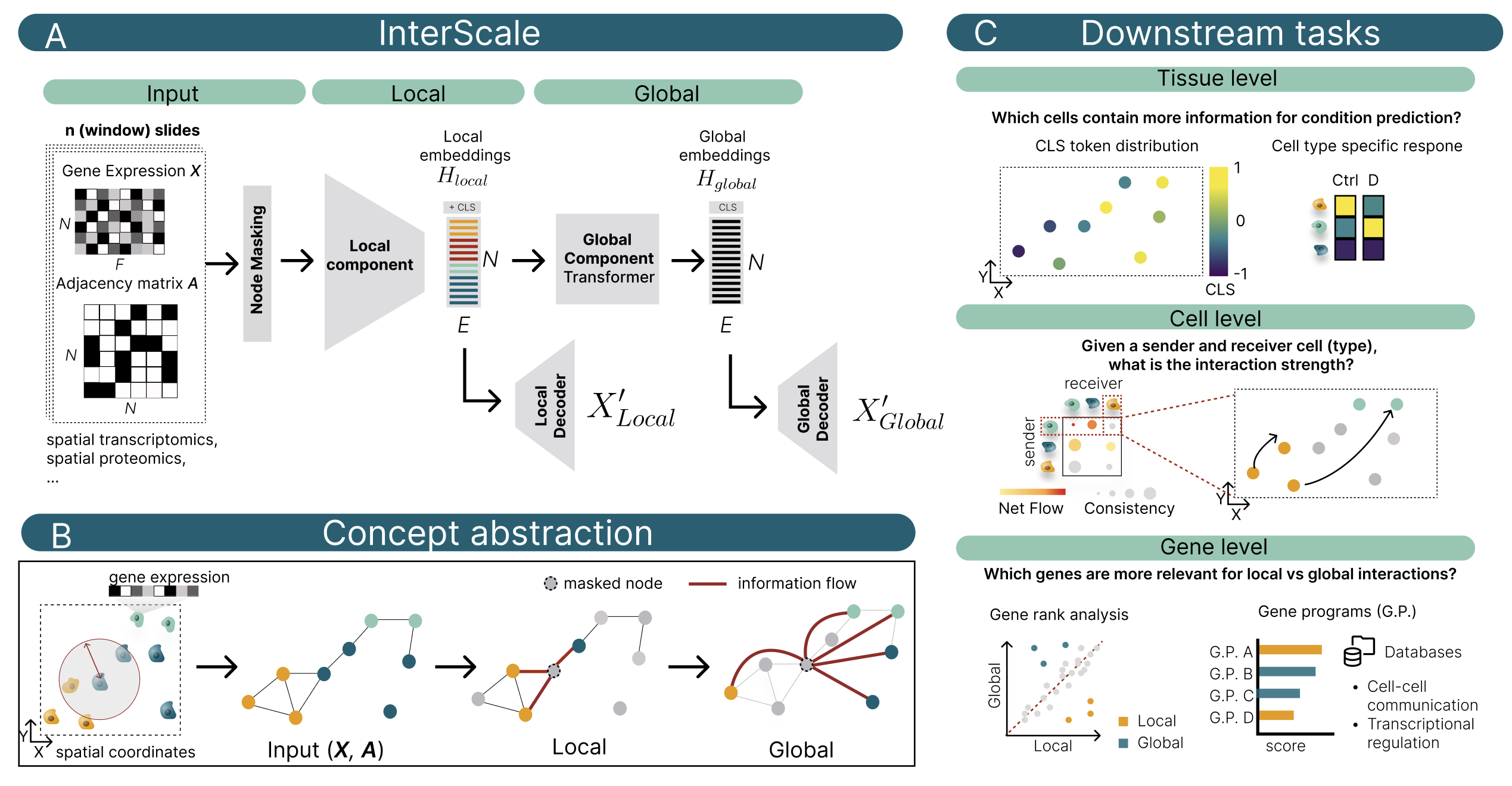
InterScale is a package for multi-scale analysis of cellular interactions in spatial transcriptomics data. It is built on top of geome (single-cell building on top of PyG), AnnData and scvi-tools.
Installation
Check out the installation guide.
Tutorials
Learn by following example application of InterScale.
API
Find a detailed description of InterScales APIs.
Release Notes
Follow the latest changes to InterScale.
Contributing
Help improve InterScale.
References
References to supporting packages and publications used in InterScale.
If you find InterScale useful for your research, please consider citing the InterScale preprint.
