Set up and download - Molecular cartography data from Legnini et al., 2023#
This tutorials shows how to set up an multiple slide molecular cartography dataset used from Legnini et al., 2023. To follow along with this and the following tutorials, please execute the following steps first:
Set up InterScale environment (see instructions in installation)
Download the sample data from the original publication from Zenodo under the accession no. 6143561
import scanpy as sc
import squidpy as sq
import numpy as np
import pandas as pd
import matplotlib.pyplot as plt
from scipy.sparse import issparse, csr_matrix
import seaborn as sns
from pathlib import Path
import warnings
warnings.filterwarnings("ignore")
from interscale import datasets as ds
from interscale.pp import compute_neighborhood_stats
Path.cwd().resolve().parent.parent
PosixPath('/Users/francesca.drummer/Documents/1_Projects/A3-InterScale/interscale')
from pathlib import Path
import sys
# Find repo root by locating paths.py
BASE_DIR_PROJECT = Path.cwd().resolve().parent.parent
sys.path.insert(0, str(BASE_DIR_PROJECT))
DATA = "legnini"
Select the sample IDs for plotting:
slide4_A2-3: Controlslide4_B2-2: SHH
sample_id = ["slide4_A2-3", "slide4_B2-2"]
Load data#
adata = ds.legnini(f"{BASE_DIR_PROJECT}/data/{DATA}")
adata
AnnData object with n_obs × n_vars = 43910 × 88
obs: 'Cell', 'Area', 'x', 'y', 'sample', 'condition', 'organoid'
var: 'gene_ids', 'feature_types'
obsm: 'spatial'
layers: 'raw'
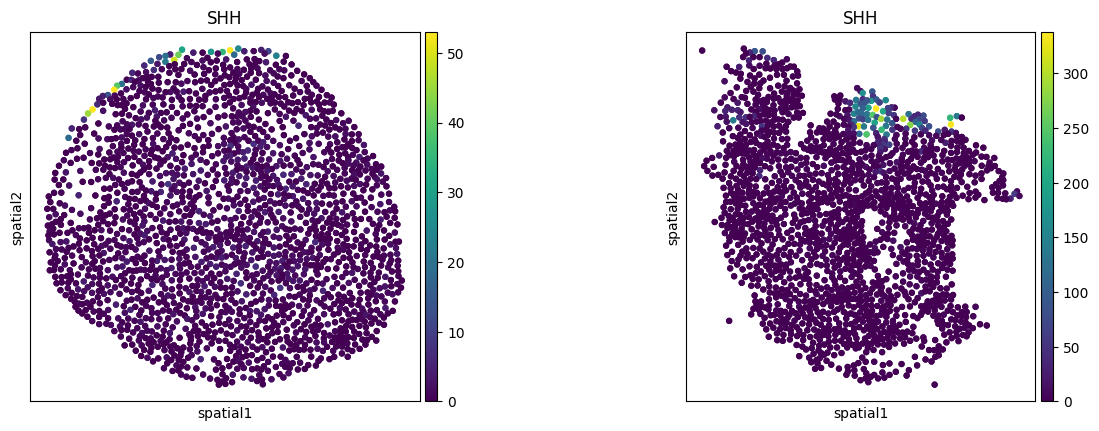
sq.pl.spatial_scatter(
adata, color=["SHH"], spatial_key="spatial", library_key="sample", library_id=sample_id, shape=None, size=50
)
print("Zero count cells: ", (adata.X.sum(1) == 0).sum())
Zero count cells: 648
np.isnan(adata.obsm["spatial"]).sum()
np.int64(296)
Remove all cells that have entry NaN in .obsm['coordinates'].
If NaN is in .obsm it leads to error when creating graph.
# Create a boolean mask for rows without NaN coordinates
valid_coords = ~np.isnan(adata.obsm["spatial"]).any(axis=1)
# Filter the AnnData object
adata = adata[valid_coords].copy()
# Verify the removal of NaN values
print(f"NaN values remaining: {np.isnan(adata.obsm['spatial']).sum()}")
NaN values remaining: 0
1. Normalization#
The data needs to be normalized for InterScale (Ideally, counts should be normalized between 0 to 3). Check if the data is already normalized:
scales_counts = sc.pp.normalize_total(adata, target_sum=None, inplace=False)
# log1p transform
adata.layers["raw"] = adata.X
adata.layers["log1p_norm"] = sc.pp.log1p(scales_counts["X"], copy=True)
# Freeman-Tukey square root transform
assert issparse(adata.X)
sqrt_X = adata.X.sqrt()
# Create a new sparse matrix for X + 1
X_plus_1 = adata.X + csr_matrix(np.ones(adata.X.shape))
# Calculate the square root of (X + 1)
sqrt_X_plus_1 = X_plus_1.sqrt()
adata.layers["norm_ftsqrt"] = sqrt_X + sqrt_X_plus_1
# shifted Logarithm
scales_counts = sc.pp.normalize_total(adata, target_sum=10000, inplace=False)
# log1p transform
adata.layers["log1p_norm"] = sc.pp.log1p(scales_counts["X"], copy=True)
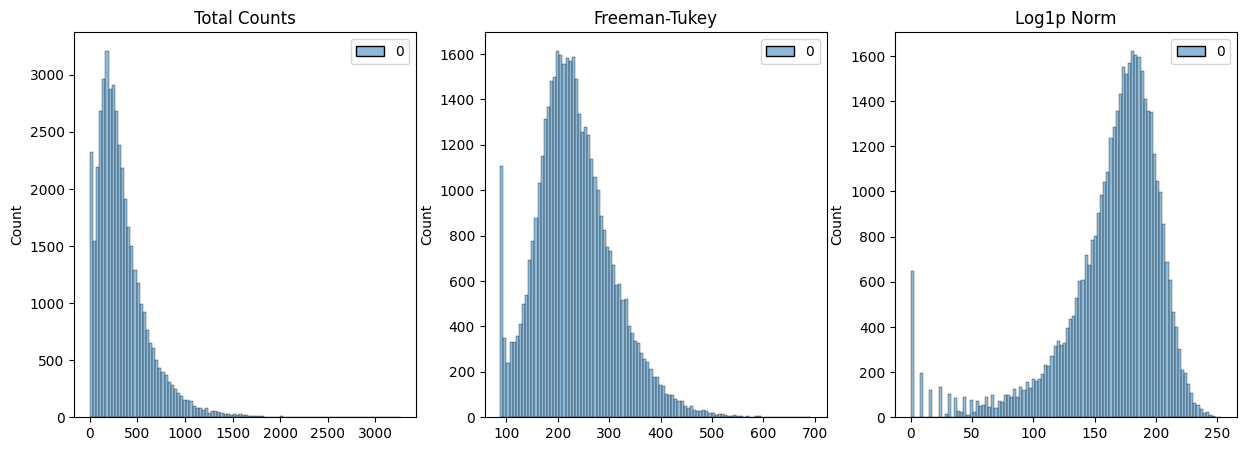
fig, axes = plt.subplots(1, 3, figsize=(15, 5))
p0 = sns.histplot(adata.layers["raw"].sum(1), bins=100, kde=False, ax=axes[0])
axes[0].set_title("Total Counts")
p1 = sns.histplot(adata.layers["norm_ftsqrt"].sum(1), bins=100, kde=False, ax=axes[1])
axes[1].set_title("Freeman-Tukey")
p2 = sns.histplot(adata.layers["log1p_norm"].sum(1), bins=100, kde=False, ax=axes[2])
axes[2].set_title("Log1p Norm")
plt.show()
print("Raw - Min: ", {adata.layers["raw"].min()}, ", Max: ", {adata.layers["raw"].max()})
print("Log1pNorm - Min: ", {adata.layers["log1p_norm"].min()}, ", Max: ", {adata.layers["log1p_norm"].max()})
print("NormTRSqrt - Min: ", {adata.layers["norm_ftsqrt"].min()}, ", Max: ", {adata.layers["norm_ftsqrt"].max()})
Raw - Min: {np.float32(0.0)} , Max: {np.float32(603.0)}
Log1pNorm - Min: {np.float32(0.0)} , Max: {np.float32(9.210441)}
NormTRSqrt - Min: {np.float64(1.0)} , Max: {np.float64(49.132470338556004)}
2. Calculate spatial connectivity matrix#
Use squidpy.gr.spatial_neighbors()) to calculate the spatial connectivity. For image-based ST it is important to set coord_type='generic'. In Squidpy, you have the option between k-nearest neighbors, delaunay and radius based neighborhood. For InterScale, we use a radius-based neighborhood to capture density information. Find the radius for which the number of connected neighbors is approximately 10-30, depending on tissue density.
stats = compute_neighborhood_stats(adata, radii=[0, 200, 300], library_key="sample")
INFO Creating graph using `generic` coordinates and `None` transform and `17` libraries.
Radius: 0, Average Neighbors: 0.00, Std Dev: 0.00
INFO Creating graph using `generic` coordinates and `None` transform and `17` libraries.
Radius: 200, Average Neighbors: 10.14, Std Dev: 2.96
INFO Creating graph using `generic` coordinates and `None` transform and `17` libraries.
Radius: 300, Average Neighbors: 23.13, Std Dev: 5.95
Make sure that obs_names are unique and convertable to string.
adata.obs_names_make_unique
<bound method AnnData.obs_names_make_unique of AnnData object with n_obs × n_vars = 43762 × 88
obs: 'Cell', 'Area', 'x', 'y', 'sample', 'condition', 'organoid'
var: 'gene_ids', 'feature_types'
uns: 'spatial_neighbors'
obsm: 'spatial'
layers: 'raw', 'log1p_norm', 'norm_ftsqrt'
obsp: 'spatial_connectivities', 'spatial_distances'>
adata.obs["obs_names"] = adata.obs_names
sq.gr.spatial_neighbors(
adata,
coord_type="generic",
library_key="sample",
radius=300,
)
INFO Creating graph using `generic` coordinates and `None` transform and `17` libraries.
# Calculate nr. of neighbors per cells
conn = adata.obsp["spatial_connectivities"]
# Print average number of connections per node
avg_connections = conn.nnz / conn.shape[0] # total connections / number of nodes
print(f"Average number of connections per node: {avg_connections:.2f}")
Average number of connections per node: 23.13
sq.pl.spatial_scatter(
adata,
color=["SHH"],
spatial_key="spatial",
library_key="sample",
library_id=sample_id,
shape=None,
connectivity_key="spatial_connectivities",
size=50,
)
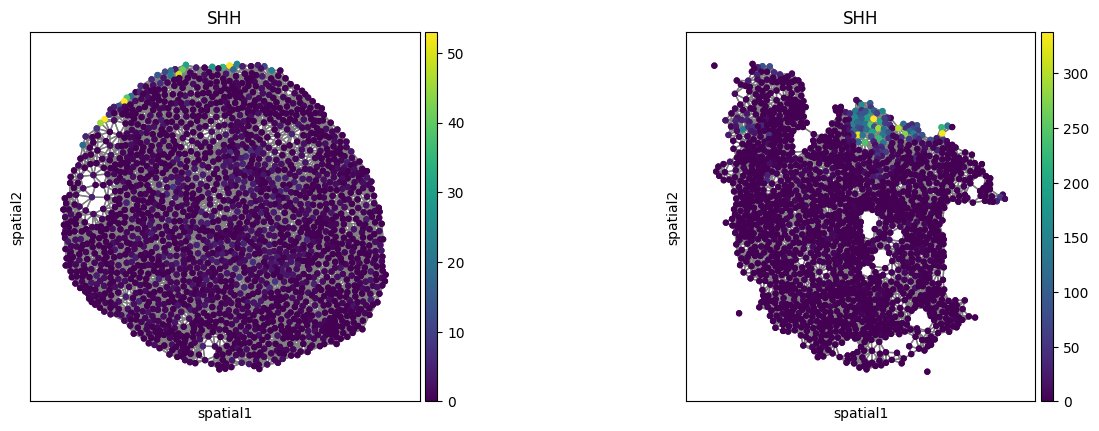
3. Optional: Calculate sliding windows#
Sliding windows are necessary in case the tissue slide contains more than 4k cells. First, check how many cells are at minimum or maximum in your dataset.
tissue_cell_number = adata.obs.groupby("sample").size()
print(
f"Nr cells per sliding window: Min: {tissue_cell_number.min()}, Max: {tissue_cell_number.max()}, Avg: {tissue_cell_number.mean()}"
)
Nr cells per sliding window: Min: 1640, Max: 4318, Avg: 2574.235294117647
In case of the Legnini data, we set the maximum number of cells to 4318.
MAX_CELLS = 4318
4. Optional: split data into train and val set#
Training the model requires a split assignment for each donor/patient/sliding window that you wanna train on.
df = adata.obs[["condition", "sample"]]
value_counts = pd.DataFrame(df.values, columns=df.columns).value_counts()
print(value_counts)
condition sample
SHH slide1_D2-2 4318
slide1_D2-3 3621
slide1_C2-2 3510
slide4_B2-1 3345
slide1_B2-1 3299
slide1_A2-1 3023
Ctrl slide1_C2-1 2870
SHH slide4_A2-1 2630
slide4_A2-2 2506
slide1_B2-3 2241
Ctrl slide4_A2-3 1984
slide1_C2-5 1958
SHH slide4_B2-2 1779
Ctrl slide1_B2-2 1701
SHH slide1_A2-2 1694
Ctrl slide4_B2-3 1643
slide1_C2-3 1640
Name: count, dtype: int64
adata.obs["split"] = "train"
# assign each one [Long-duration, ND, Onset]
adata.obs["split"][
adata.obs["sample"].isin(["slide1_B2-1", "slide1_B2-3", "slide1_A2-2", "slide1_C2-3", "slide4_B2-3"])
] = "val"
adata.obs["split"][adata.obs["sample"].isin(sample_id)] = "test"
df = adata.obs[["split", "sample", "condition"]]
value_counts = pd.DataFrame(df.values, columns=df.columns).value_counts()
print(value_counts)
split sample condition
train slide1_D2-2 SHH 4318
slide1_D2-3 SHH 3621
slide1_C2-2 SHH 3510
slide4_B2-1 SHH 3345
val slide1_B2-1 SHH 3299
train slide1_A2-1 SHH 3023
slide1_C2-1 Ctrl 2870
slide4_A2-1 SHH 2630
slide4_A2-2 SHH 2506
val slide1_B2-3 SHH 2241
test slide4_A2-3 Ctrl 1984
train slide1_C2-5 Ctrl 1958
test slide4_B2-2 SHH 1779
train slide1_B2-2 Ctrl 1701
val slide1_A2-2 SHH 1694
slide4_B2-3 Ctrl 1643
slide1_C2-3 Ctrl 1640
Name: count, dtype: int64
Save adata object#
Save the prepared adata object such that it can be loaded for the model training.
adata.write(f"{BASE_DIR_PROJECT}/data/{DATA}_pp.h5ad")
Prepare config file#
Duplicate the minum requirement config file from /interscale/config_files/legnini_example.yaml and add the necessary specification for the Visum data:
model:
local_component:
name: GCN
global_component:
name: self-attn-transformer
max_seq_len: 2418
save: /path/to/save/model/
dataset:
h5ad_data: legnini22.h5ad
name: legnini23
sample_key: ['sample']
spatial_neigbors_kwargs:
coord_type: generic
library_key: sample
radius: 200
Save the config file as .yaml and proceed to training (either interactively in jupyter notebook or by running a script).
