Model#
InterScale is a model descigned for spatial transcpriptomics analysis. It provides, 1) local and global embeddings for gene level analysis and 2) attention matrix for cell-to-cell analysis.
Overview#
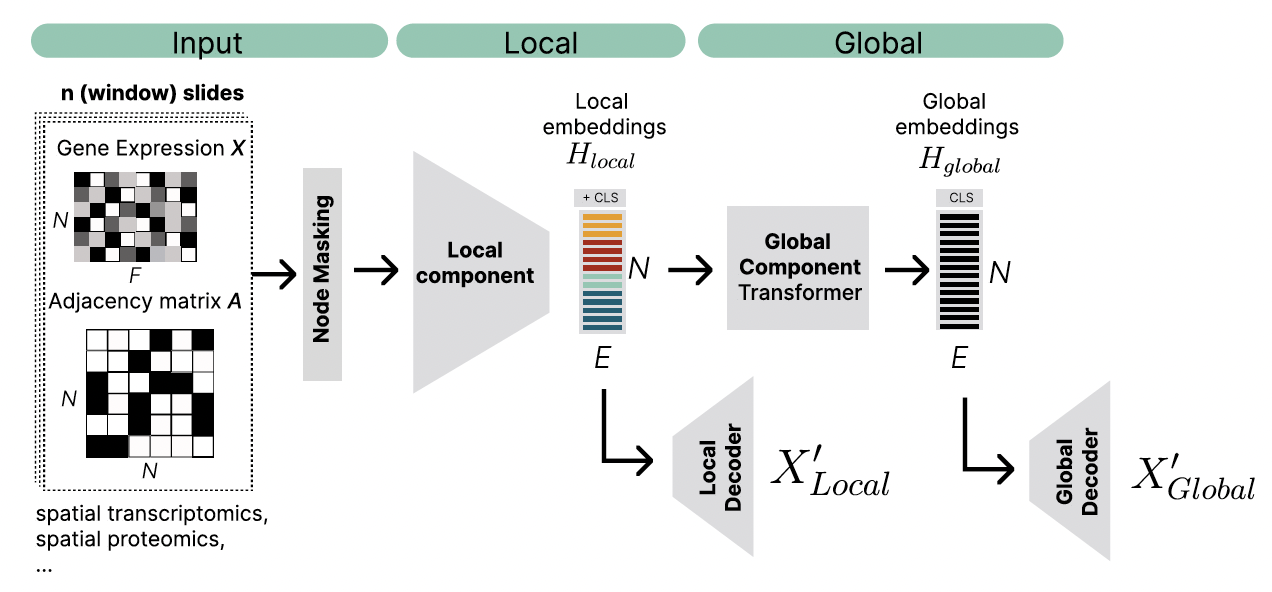
InterScale is a two component model. The local model learns cell representation of a local, spatial neighborhood and the global compponent learns tissue wide interactions between these neighborhoods.
InterScale model#
This is the main model class that can be used to define, train, and evaluate the model on an anndata. InterScale’s model uses module to initialize the local and global components in the model (see below).
Combined model with both local and global components. |
|
Global model with only global component. |
|
Local model with only local component. |
InterScale module#
InterScale is built from composable local and global modules. The combined modules wire them together into a full model. The base classes (LocalModule, GlobalModule) define the interface — subclass either to implement a custom architecture and pass it to any model class.
Combined modules#
Two combined modules are available. DualDecoderCombinedModule is the default and trains a separate decoder for each scale. CombinedModule uses a single shared (global) decoder.
Combined module with decoders for both local and global modules. |
|
Combined module with single decoder for both local and global modules. |
Local modules#
LocalModule is the base class. Four ready-made implementations are provided; subclass LocalModule to define your own.
Graph Isomorphism Network (local module) - builds on PyTorch Geometric implementation. |
|
Graph Convolutional Network (local module) - builds on PyTorch Geometric implementation. |
|
scVI Encoder (local module) - builds on scvi-tools implementation. |
|
Module for using frozen, pre-computed embeddings. |
Global modules#
GlobalModule is the base class. The default implementation is a transformer encoder with self-attention relevance hooks; subclass GlobalModule to define your own.
Sequence of: Dropout → Layer Norm → FC → nonlinearity → Dropout → FC → Dropout → Layer Norm + residual connections |
Usage example#
import scanpy as sc
from interscale
from interscale.tl import prepare_geome_dataset
from interscale.geome_dataloader import GraphAnnDataModule
# Load your model and training configurations
cfg = load_config(cfg_path)
# Load your data
adata = ad.read_h5ad("your_data.h5ad")
# Setup anndata
interscale.model.CombinedModel._setup_anndata(
adata = adata,
prediction_task = PREDICTION_TASK,
layer_key = "norm",
sample_key_list = ["sample"],
prediction_obs = prediction_obs
)
# Initialize the model
model = interscale.model.CombinedModel(
adata,
cfg = cfg
)
pyg_data_list, _ = prepare_geome_dataset(adata, cfg)
dm = GraphAnnDataModule(datas=pyg_data_list,
num_workers=1,
batch_size=int(cfg.dataset.batch_size),
pct_mask_nodes=cfg.dataset.pct_mask_nodes,
learning_type="node")
# Train the model
model.train(max_epochs = 20,
datamodule = dm,
early_stopping = True,
batch_size = int(cfg.dataset.batch_size),
train_size = float(cfg.dataset.train_size),
validation_size = float(cfg.dataset.val_size),
wandb_use = False)
# Get model output
result = model.get_model_output(adata)
# Please check tutorials for more details and downstream steps
